<!-- bara: put body text here -->
<u>Section list:</u>
<ul>
<li><a href='#General'>General</a></li>
<li><a href='#Polymorphism'>Polymorphism</a></li>
<li><a href='#Expression in tissues (from GTEx)'>Expression in tissues (from GTEx)</a></li>
</ul>
------
<h2 id='General'>General</h2>
| | |
|--|--|
| <b>Gene ensembl <br><i>Link to the corresponding Ensembl site</i></b> | <a href="http://www.ensembl.org/Multi/Search/Results?q=ENSG00000269693;site=ensembl~">ENSG00000269693</a> |
| <b>Age (years) <br><i>Estimated age of the KRAB-ZNF as in <a href="https://pubmed.ncbi.nlm.nih.gov/28273063/">Imbeault et al</a></i></b> | 81.3 |
| <b>Age (species) <br><i>The name of the species where the KRAB-ZNF was first observed as in <a href="https://pubmed.ncbi.nlm.nih.gov/28273063/">Imbeault et al</a></i></b> | Sunda flying lemur |
| <b>\#peaks in exo <br><i>Number of peaks found in the ChIP-exo dataset</i></b> | |
| <b>Incomplete KRAB domain</b> | |
| <b>Interacts with K1</b> | Not in database |
| <b>Interacts with</b> | Not in database |
| <b>ORF sequence</b> | |
| <b>Zinc fingers</b> | 1 |
<div class="adroite"><a href="#"><b>back to top</b></a></div>
------
<h2 id='Polymorphism'>Polymorphism</h2>
<table>
<tr><th>gnomAD genome</th><th>gnomAD exome</th></tr>
<tr><td><table></table>
| Region | #miss.SNP x Kb |
|--|--|
| Exon | 172.272 |
| Exon (no domain, nor Zfinger) | 172.272 |
| Krab domain | NA |
| Zinc finger | NA |
| Region | #miss.SNP |
|--|--|
| Zinc finger | NA |
| C2H2 | NA |
| Zinc finger print | NA |
<input type="button" id="button" style="float:left" value="[+] Help" onclick="afficher()" /><br>
<form id="doc" style="display: none;">
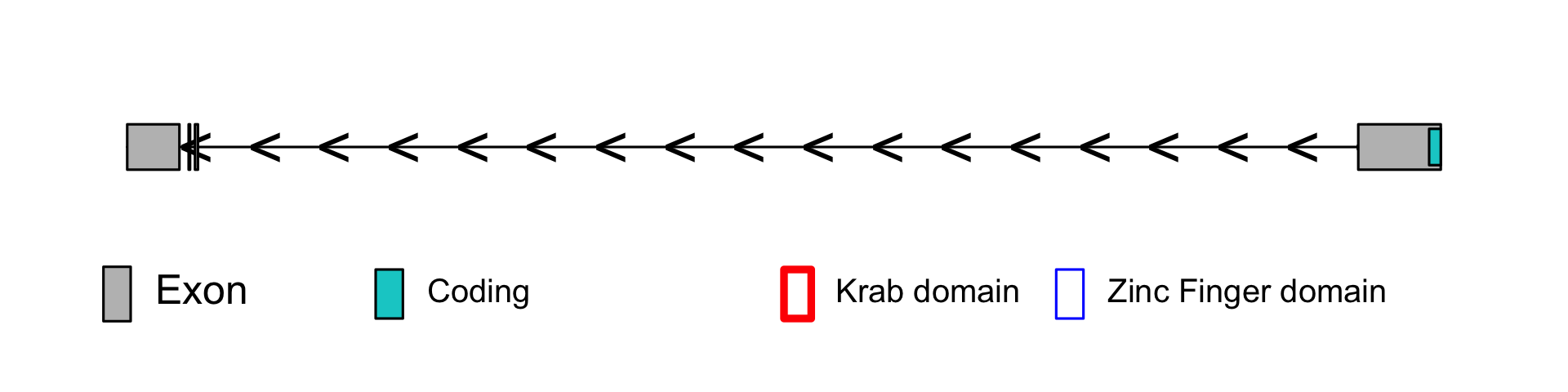
<br>Gene view with all variants found at the gene of interest (canonical transcript).<br><ul><li> blue dots: synonymous & noncoding variants.</li><li> red dots: missense variants.</li><li> orange dots: splice variants.</li><li> green dots: Loss of function variants (frameshift, stop_lost or stop_gained mutation).</li><li> GERP score: a conservation score widely used to look at constraint across genes and non-coding regions, and also used in the best pathogenic variant scoring softwares. A score above 2 is considered conserved.</li><li> Green boxes on the genes: the exons.</li><li> Red boxes: the KRAB domain.</li><li> Blue boxes: the zinc finger domains.</li><li> Pink boxes: degenerate zinc finger domains.</li></ul>
</form><br>
<a href='../MAF_gnomAD/CTD-2192J16.20.pdf'>
<img src="../MAF_gnomAD/MAF_gnomAD/CTD-2192J16.20.png" width="400" height="200" />
</a>
<input type="button" id="button_2" style="float:left" value="[+] Help" onclick="afficher_2()" /><br>
<form id="doc_2" style="display: none;">
<br>Zoomed plot of variants in each zinc finger domain with the specific amino acids plotted.<ul><li>blue dots: synonymous & noncoding variants.</li><li>red dots: missense variants.</li><li>orange dots: splice variants.</li><li>green dots: Loss of function variants (frameshift, stop_lost or stop_gained mutation).</li><li> Blue boxes: The positions of the DNA contacting amino acids</li><li> Red boxes: The Cytidine and Histidine amino acid positions</li></ul>
</form><br>
<a href='../Zfinger_gnomAD/CTD-2192J16.20.pdf'>
<img src="../Zfinger_gnomAD/Zfinger_gnomAD/CTD-2192J16.20.png" width="400" height="200" />
</a>
</td><td><table></table>
| Region | #miss.SNP x Kb |
|--|--|
| Exon | 218.03 |
| Exon (no domain, nor Zfinger) | 218.03 |
| Krab domain | NA |
| Zinc finger | NA |
| Region | #miss.SNP |
|--|--|
| Zinc finger | NA |
| C2H2 | NA |
| Zinc finger print | NA |
<input type="button" id="button1" style="float:left" value="[+] Help" onclick="afficher1()" /><br>
<form id="doc1" style="display: none;">
<br>Gene view with all variants found at the gene of interest (canonical transcript).<br><ul><li> blue dots: synonymous & noncoding variants.</li><li> red dots: missense variants.</li><li> orange dots: splice variants.</li><li> green dots: Loss of function variants (frameshift, stop_lost or stop_gained mutation).</li><li> GERP score: a conservation score widely used to look at constraint across genes and non-coding regions, and also used in the best pathogenic variant scoring softwares. A score above 2 is considered conserved.</li><li> Green boxes on the genes: the exons.</li><li> Red boxes: the KRAB domain.</li><li> Blue boxes: the zinc finger domains.</li><li> Pink boxes: degenerate zinc finger domains.</li></ul>
</form><br>
<a href='../MAF_gnomAD_exome/CTD-2192J16.20.pdf'>
<img src="../MAF_gnomAD_exome/MAF_gnomAD_exome/CTD-2192J16.20.png" width="400" height="200" />
</a>
<input type="button" id="button2" style="float:left" value="[+] Help" onclick="afficher2()" /><br>
<form id="doc2" style="display: none;">
<br>Zoomed plot of variants in each zinc finger domain with the specific amino acids plotted.<ul><li>blue dots: synonymous & noncoding variants.</li><li>red dots: missense variants.</li><li>orange dots: splice variants.</li><li>green dots: Loss of function variants (frameshift, stop_lost or stop_gained mutation).</li><li> Blue boxes: The positions of the DNA contacting amino acids</li><li> Red boxes: The Cytidine and Histidine amino acid positions</li></ul>
</form><br>
<a href='../Zfinger_gnomAD_exome/CTD-2192J16.20.pdf'>
<img src="../Zfinger_gnomAD_exome/Zfinger_gnomAD_exome/CTD-2192J16.20.png" width="400" height="200" />
</a>
</td></tr>
</table>
<div class="adroite"><a href="#"><b>back to top</b></a></div>
------
<h2 id='Expression in tissues (from GTEx)'>Expression in tissues (from GTEx)</h2>
<input type="button" id="GTX_button" style="float:left" value="Open Expression Information from GTEx Website" onclick="document.location.href='https://www.gtexportal.org/home/gene/CTD-2192J16.20'" /><br><br>
<div class="adroite"><a href="#"><b>back to top</b></a></div>
<!-- bara: end of place to put text -->